num = 37
if num > 100:
print('greater')
else:
print('not greater')
print('done')not greater
doneAt the end of this class, you will be able to:
In our last lesson, we discovered something suspicious was going on in our inflammation data by drawing some plots. How can we use Python to automatically recognize the different features we saw, and take a different action for each? In this lesson, we’ll learn how to write code that runs only when certain conditions are true.
Conditionals allows you to make decisions in your code based on certain conditions.
if something is true:
do task a
otherwise:
do task bFor example we can ask Python to take different actions, depending on a condition, with an if statement:
num = 37
if num > 100:
print('greater')
else:
print('not greater')
print('done')not greater
doneIf the test that follows the if statement is true, the body of the if (i.e., the set of lines indented underneath it) is executed, and “greater” is printed. If the test is false, the body of the else is executed instead, and “not greater” is printed. Only one or the other is ever executed before continuing on with program execution to print “done”.
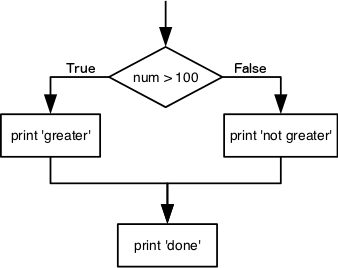
Conditional statements don’t have to include an else. If there isn’t one, Python simply does nothing if the test is false:
num = 53
print('before conditional...')
if num > 100:
print(num, 'is greater than 100')
print('...after conditional')before conditional...
...after conditionalHere we used a comparison operator (>) to compare the value of num to 100. Here are other comparison operators:
| Operator | Name |
|---|---|
== |
Equal |
!= |
Not equal |
> |
Greater than |
>= |
Greater than or equal to |
< |
Less than |
<= |
Less than or equal to |
# Example
2 == 1 + 1TrueNote that to test for equality we use a double equals sign == rather than a single equals sign = which is used to assign values.
You should never use equalty operators (==or !=) with floats or complex values.
# Example
2.1 + 3.2 == 5.3FalseThis is a floating point arithmetic problem seen in other programming languages. It is due to the difficulty of having a fixed number of binary digits (bits) to accurately represent some decimal number. This leads to small rounding errors in calculations.
2.1 + 3.2 5.300000000000001If you need to use equalty operators, do it with a degree of freedom:
tol = 1e-6 ; abs((2.1 + 3.2) - 5.3) < tolTrueThe result of a comparison operator is a boolean value (True or False), which can be used in an if statement to control the flow of the program.
Booleans represent one of two values: True or False. When you compare two values, the expression is evaluated and Python returns the Boolean answer:
num = 37
print(num > 100)FalseThere are also membership operators (in and not in). They are used to test if a value is present in a sequence (like a list or a string):
| Operator | Description |
|---|---|
in |
Returns True if a sequence with the specified value is present in the object |
not in |
Returns True if a sequence with the specified value is not present in the object |
seq = ['ATGAAGGGTCCAAAA', 'AGTCCCCGTATGAT', 'ACCT', 'ACCT']
print('ACCT' in seq)
print('G' in seq[-1])True
Falseelif statementsWe can also chain several tests together using elif which is short for “else if”. The elif keyword is Python’s way of saying “if the previous conditions were not true, then try this condition”. The following Python code uses elif to print the sign of a number.
num = -3
if num > 0:
print(num, 'is positive')
elif num == 0:
print(num, 'is zero')
else:
print(num, 'is negative')-3 is negativeWe can also combine tests using logical operators and, or and not to create more complex conditions.
Logical operators are used to combine conditional statements:
| Operator | Description |
|---|---|
and |
Returns True if both statements are true |
or |
Returns True if one of the statements is true |
not |
Reverse the result, returns False if the result is true |
Result of the and oeprator:
| Boolean 1 | Boolean 2 |
|---|---|
True |
True |
True |
False |
False |
True |
False |
False |
# Example
False and False, False and True, True and False, True and True(False, False, False, True)Result of the or oeprator:
| Boolean 1 | Boolean 2 |
|---|---|
True |
True |
True |
False |
False |
True |
False |
False |
# Example
False or False, False or True, True or False, True or True(False, True, True, True)# Example
True or not TrueTrueif (1 > 0) and (-1 >= 0):
print('both parts are true')
else:
print('at least one part is false')at least one part is falseif (1 < 0) or (1 >= 0):
print('at least one test is true')at least one test is trueNow that we’ve seen how conditionals work, we can use them to check for the suspicious features we saw in our inflammation data.
From the first couple of plots, we saw that maximum daily inflammation exhibits a strange behavior and raises one unit a day. Wouldn’t it be a good idea to detect such behavior and report it as suspicious? Let’s do that! However, instead of checking every single day of the study, let’s merely check if maximum inflammation in the beginning (day 0) and in the middle (day 20) of the study are equal to the corresponding day numbers.
import pandas as pd
data = pd.read_csv('data/inflammation-01.csv', index_col=0)
max_inflammation_0 = data.max(axis=0).iloc[0]
max_inflammation_20 = data.max(axis=0).iloc[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')Suspicious looking maxima!We also saw a different problem in the third dataset; the minima per day were all zero (looks like a healthy person snuck into our study). And if neither of these conditions are true, we can use else to give the all-clear:
data = pd.read_csv('data/inflammation-01.csv', index_col=0)
max_inflammation_0 = data.max(axis=0).iloc[0]
max_inflammation_20 = data.max(axis=0).iloc[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
elif data.min(axis=0).sum() == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')Suspicious looking maxima!With another dataset:
data = pd.read_csv('data/inflammation-03.csv', index_col=0)
max_inflammation_0 = data.max(axis=0).iloc[0]
max_inflammation_20 = data.max(axis=0).iloc[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
elif data.min(axis=0).sum() == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')Minima add up to zero!In this way, we have asked Python to do something different depending on the condition of our data. Here we printed messages in all cases, but we could also imagine not using the else catch-all so that messages are only printed when something is wrong, freeing us from having to manually examine every plot for features we’ve seen before.
With gene1_expression and gene2_expression given, are these 2 codes equivalent?
# Code A
if gene1_expression > gene2_expression:
print("Gene 1 has higher expression level.")
elif gene1_expression < gene2_expression:
print("Gene 2 has higher expression level.")
else:
print("Gene 1 and Gene 2 have the same expression level.")# Code B
if gene1_expression > gene2_expression:
print("Gene 1 has higher expression level.")
else:
if gene1_expression < gene2_expression:
print("Gene 2 has higher expression level.")
else:
print("Gene 1 and Gene 2 have the same expression level.")Are these two codes equivalent?
# Code A
if "ATG" in dna_sequence:
print("Start codon found.")
elif "TAG" in dna_sequence:
print("Stop codon found.")
else:
print("No interesting codon found.") # Code B
if "ATG" in dna_sequence:
print("Start codon found.")
if "TAG" in dna_sequence:
print("Stop codon found.")
else:
print("No interesting codon found.") This code is incorrect, can you tell why?
x = 7
if x > 5:
print("x is greater than 5")
if x > 10:
print("x is greater than 10")
elif x = 10:
print("x equals 10")
else:
print("x is less than 10") Instead of just printing if a file falls into one or the other category, we want to save in lists the name of the files that have suspicious maximum trend, a suspicious minimum trend and the ones that seem normal at the moment.
Modify the following code:
data = pd.read_csv('data/inflammation-03.csv', index_col=0)
max_inflammation_0 = data.max(axis=0).iloc[0]
max_inflammation_20 = data.max(axis=0).iloc[20]
if max_inflammation_0 == 0 and max_inflammation_20 == 20:
print('Suspicious looking maxima!')
elif data.min(axis=0).sum() == 0:
print('Minima add up to zero!')
else:
print('Seems OK!')Minima add up to zero!You will need to loop through all the files in the data folder, and create three lists maxima_suspicious_files, minima_suspicious_files and normal_files to save the names of the files that fall into each category.
Remember to make use of the function list.append(element) to add an element to a list.
In the last exercise, we created three lists to store the names of the files that have suspicious maximum trend, a suspicious minimum trend and the ones that seem normal at the moment. We could also have stored this information in a dictionary, which is a data structure that allows us to store key: value pairs.
A dictionary is a collection which is ordered (as of Python >= 3.7), changeable and does not allow duplicates keys. Dictionaries are written with curly brackets {}, with keys and values. Keys must be unique and of an immutable type (like strings or numbers), while values can be of any type (like strings, numbers, lists, dictionnaries…) and can be duplicated.
organism1_genes = {
#key: value;
'BRCA1': 'DNA repair',
'TP53': 'Tumor suppressor',
'EGFR': 'Cell growth',
'MYC': 'Regulation of gene expression'
}Dictionary items can be referred to by using the key name.
organism1_genes["BRCA1"]'DNA repair'Dictionaries have specific methods. Here are a few:
| Method | Description |
|---|---|
.items() |
Returns a list containing a tuple for each key value pair |
.keys() |
Returns a list containing the dictionary’s keys |
.values() |
Returns a list of all the values in the dictionary |
.pop() |
Removes the element with the specified key |
.get() |
Returns the value of the specified key |
Modify the previous code so that instead of having 3 lists maxima_suspicious_files, minima_suspicious_files and normal_files you have only one dictionnary, called classify_files. The values of the dictionary should be a list containing the name of the files, and the keys should be one of the following strings: "suspicious maxima", "suspicious minima" or "normal".
You can initialize the dictionnary like so:
categorize_files = {
"suspicious maxima": [],
"suspicious minima": [],
"normal": []
}From the dictionary organism1_genes created as example, get the value of the key BRCA1. If the key does not exist, return Unknown by default. Try your code before and after removing the BRCA1 key:value pair.
Check the help of get by running help(dict.get).
We have created a workflow to check if our data looks suspicious. In python we can wrap this workflow into a function, which is a reusable block of code that performs a specific task. Functions allow us to organize our code and make it more modular and easier to read.
For example, instead of reading a whole block of code, we could create a function called check_suspicious_data() that takes a filename as input and returns whether the data in the file is suspicious or not. But first let’s learn how to create a function in Python.
A function usually takes some data as input (parameters that are required or optional), and usually returns an output (that can be of any type).
We already learned how to run a predefined function in the last lesson. You need to write its name followed by parenthesis. Parameters are added inside the parenthesis as follow:
# round(number, ndigits=None)
x = round(number = 5.76543, ndigits = 2)
print(x)5.77To get more information about a function, use the help() function.
We will now learn how to create our own function.
Imagine that we want to convert a temperature in Fahrenheit and converting it to Celsius. We could write:
fahrenheit_val = 99
celsius_val = ((fahrenheit_val - 32) * (5/9))and for a second number we could just copy the line and rename the variables:
fahrenheit_val = 99
celsius_val = ((fahrenheit_val - 32) * (5/9))
fahrenheit_val2 = 43
celsius_val2 = ((fahrenheit_val2 - 32) * (5/9))But we would be in trouble as soon as we had to do this more than a couple times. Cutting and pasting it is going to make our code get very long and very repetitive, very quickly. We’d like a way to package our code so that it is easier to reuse, a shorthand way of re-executing longer pieces of code. This is what functions are for.
In python, a function is declared with the keyword def followed by its name, and the arguments inside parenthesis. The next block of code, corresponding to the content of the function, must be indented. The output is defined by the return keyword.
def explicit_fahr_to_celsius(temp):
# Assign the converted value to a variable
converted = ((temp - 32) * (5/9))
# Return the value of the new variable
return converted
def fahr_to_celsius(temp):
# Return converted value more efficiently using the return
# function without creating a new variable. This code does
# the same thing as the previous function but it is more explicit
# in explaining how the return command works.
return ((temp - 32) * (5/9))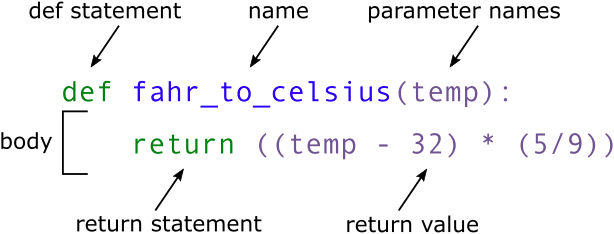
When we call the function, the values we pass to it are assigned to those variables so that we can use them inside the function. Inside the function, we use a return statement to send a result back to whoever asked for it. Let’s try running our function:
converted_temp = fahr_to_celsius(32)
print(converted_temp)
# or even,
print('freezing point of water:', fahr_to_celsius(32), 'C')
print('boiling point of water:', fahr_to_celsius(212), 'C')0.0
freezing point of water: 0.0 C
boiling point of water: 100.0 CWe’ve successfully called the function that we defined, and we have access to the value that we returned.
You can also add a description of the function directly after the function definition. It is the message that will be shown when running help(). As it can be along text over multiple lines, it is common to put it inside triple quotes """.
def fahr_to_celsius(temp, inverted = False):
"""Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
"""
return ((temp - 32) * (5/9))
help(fahr_to_celsius)Help on function fahr_to_celsius in module __main__:
fahr_to_celsius(temp, inverted=False)
Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
You can have several arguments. They can be mandatory or optional. To make them optional, they need to have a default value assigned inside the function definition, like so:
def fahr_to_celsius(temp, inverted = False):
"""Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
"""
if inverted:
new_temp = ((temp * (9/5)) + 32)
else:
new_temp = ((temp - 32) * (5/9))
return new_temp
# Or more efficiently:
def fahr_to_celsius(temp, inverted = False):
"""Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
"""
if inverted:
return ((temp * (9/5)) + 32)
else:
return ((temp - 32) * (5/9))
help(fahr_to_celsius)Help on function fahr_to_celsius in module __main__:
fahr_to_celsius(temp, inverted=False)
Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
The return keyword is present twice, but the function will stop as soon as it reaches the first return statement, so if inverted is True, the second return statement will never be reached.
The input temp is mandatory but inverted is optional as it has a default value of False. If we call the function without providing a value for inverted, it will be set to False by default.
fahr_to_celsius(32)0.0fahr_to_celsius(inverted = True)--------------------------------------------------------------------------- TypeError Traceback (most recent call last) Cell In[44], line 1 ----> 1 fahr_to_celsius(inverted = True) TypeError: fahr_to_celsius() missing 1 required positional argument: 'temp'
Reminder: if you provide the parameters in the exact same order as they are defined, you don’t have to name them. If you name the parameters you can switch their order. As good practice, put all required parameters first.
fahr_to_celsius(inverted = True, temp = 100)212.0fahr_to_celsius(100, True)212.0If no return statement is given, then no output will be returned, but the function will be run.
def fahr_to_celsius(temp, inverted = False):
"""Returns the temperature in Celsius given a temperature in Fahrenheit,
or the inverse (Celsius to Fahrenheit) if inversted is True.
Parameters:
temp (float): Temperature in Fahrenheit.
inverted (bool, optional): Whether to convert from Celsius to Fahrenheit (True) or Fahrenheit to Celsius (False).
"""
print("We are inside the function, the value of temp is:", temp)
if inverted:
new_temp = ((temp * (9/5)) + 32)
else:
new_temp = ((temp - 32) * (5/9))print(fahr_to_celsius(32))We are inside the function, the value of temp is: 32
NoneThe output can be of any type. If you have a lot of things to return, you might want to return a list.
def multiple_of_3(list_of_numbers):
"""Returns the number that are multiple of 3."""
multiples = []
for num in list_of_numbers:
if num % 3 == 0:
# num % 3 == 0 means that the remainder
# of num divided by 3 is 0,
# which means that num is a multiple of 3
multiples.append(num)
return multiples
multiple_of_3(range(1, 20, 2))[3, 9, 15]This could be written as a one-liner.
def multiple_of_3(list_of_numbers):
"""Returns the number that are multiple of 3."""
multiples = [num for num in list_of_numbers if num % 3 == 0]
return multiples
multiple_of_3(range(1, 20, 2))[3, 9, 15]In composing our temperature conversion function, we sometimes created a variable inside of the function, new_temp. We refer to such variables as local variables because they no longer exist once the function is done executing. If we try to access their values outside of the function, we will encounter an error:
print(new_temp)--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[51], line 1 ----> 1 print(new_temp) NameError: name 'new_temp' is not defined
If you want to reuse the converted temperature having calculated it with fahr_to_celsius, you can store the result of the function call in a variable:
converted_temp = fahr_to_celsius(32)
print("Converted temp is:", converted_temp)We are inside the function, the value of temp is: 32
Converted temp is: NoneThe variable converted_temp, being defined outside any function, is said to be global.
Interestingly, inside a function, one can read the value of such global variables:
def print_temperatures():
print('temperature in Fahrenheit was:', temp_fahr)
print('temperature in Celsius was:', temp_celsius)
temp_fahr = 212.0
temp_celsius = fahr_to_celsius(212.0)
print_temperatures()We are inside the function, the value of temp is: 212.0
temperature in Fahrenheit was: 212.0
temperature in Celsius was: NoneIt is not recommended to modify the value of a global variable inside a function, as it can lead to unexpected behavior and make the code harder to debug. If you need to modify a global variable, it is better to return the modified value from the function and assign it to the global variable outside of the function.
Now that we know how to wrap bits of code up in functions, we can make our inflammation analysis easier to read and easier to reuse. First, let’s make a visualize() function that generates our plots:
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
def visualize(filename):
"""Visualizes the average, maximum and minimum inflammation per day for a given file.
Parameters:
filename (str): The path of the file to visualize.
"""
data = pd.read_csv(filename, index_col=0)
fig = plt.figure()
axes1 = fig.add_subplot(1, 3, 1)
axes2 = fig.add_subplot(1, 3, 2)
axes3 = fig.add_subplot(1, 3, 3)
axes1.set_ylabel('average')
axes1.set_xlabel('Days')
axes1.set_xticks([])
axes1.plot(data.mean(axis=0))
axes2.set_ylabel('max')
axes2.set_xlabel('Days')
axes2.set_xticks([])
axes2.plot(data.max(axis=0))
axes3.set_ylabel('min')
axes3.plot(data.min(axis=0))
axes3.set_xlabel('Days')
axes3.set_xticks([])
fig.tight_layout()
plt.show()
visualize("data/inflammation-01.csv")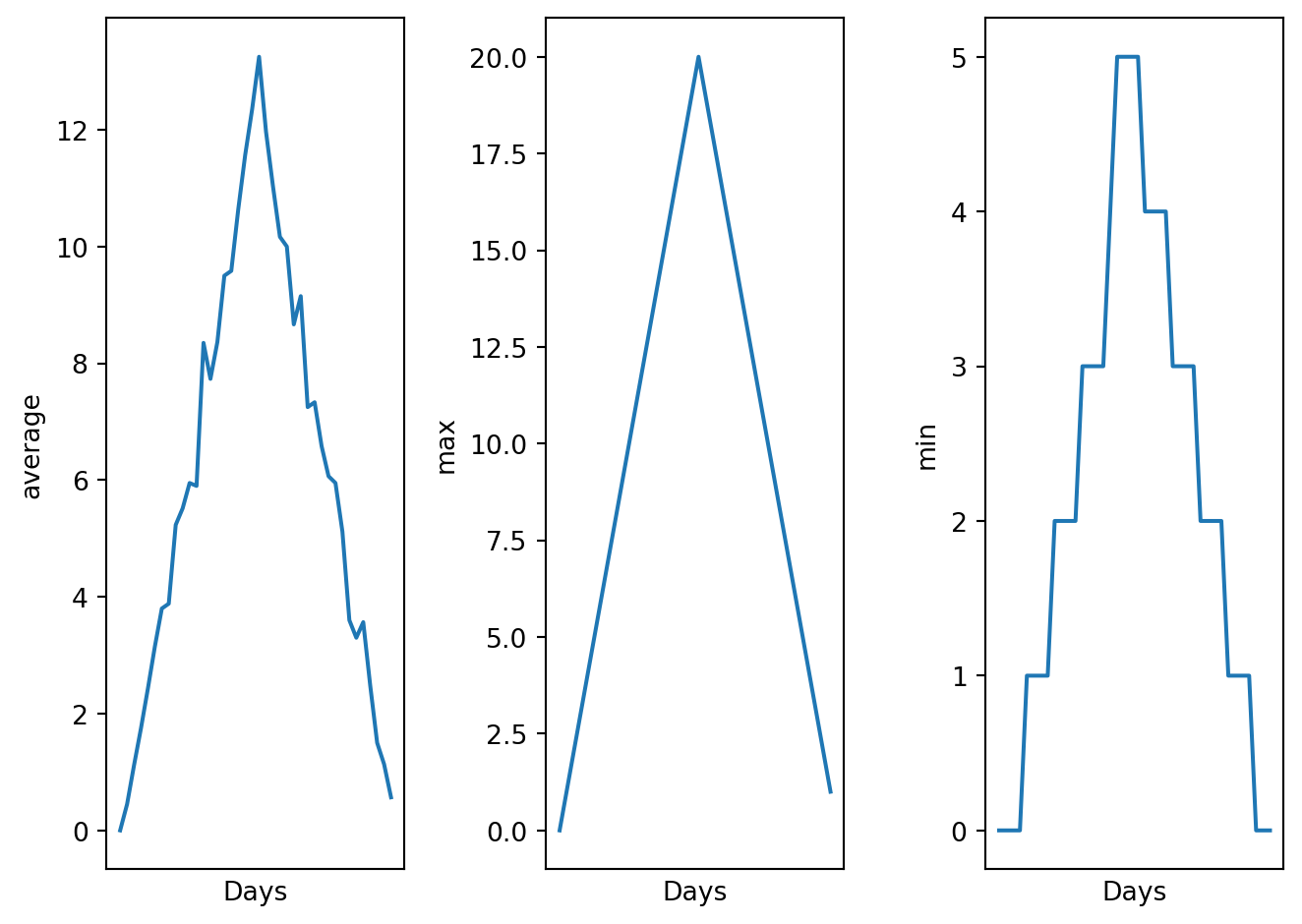
We did not use the return keyword. In Python, functions are not required to include a return statement and can be used for the sole purpose of grouping together pieces of code that conceptually do one thing. In such cases, function names usually describe what they do, e.g. visualize. This is a common practice in Python, and it is not considered bad style. In fact, it can make the code more readable and easier to understand.
Create a function that prints a message if the data in a file looks suspicious, using the same criteria as before (suspicious looking maxima and minima that add up to zero). No output needs to be returned, just print the message. It needs to also print the name of the file that is being checked.
Use it on one of the inflammation files, and then on all of them using a loop.
Add an optional parameter that asks the user to visulize the data if it has a suspicious maxima. If the parameter is set to True, the function should call the visualize() function we created earlier to show the mean, max and min plots of the data.
Sooner or later we will want to use our program in a pipeline or run it in a shell script to process thousands of data files. In order to do that in an efficient way, we need to make our programs work like other Unix command-line tools. For example, we may want a program that reads a dataset and prints the average inflammation per patient.
If you remember from the first lesson, it can be used either interactively in the terminal, or by writing a shell script (a file that contains shell commands) and running it.
In this lesson we are switching from typing commands in a Python interpreter to typing commands in a shell terminal window (such as bash). When you see a $ in front of a command that tells you to run that command in the shell rather than the Python interpreter.
For this we need to create a Python script, which is a file that contains Python code. We can create a Python script using any text editor. The file should have a .py extension to indicate that it is a Python script.
Open a new file in your text editor, and write the following code:
#!/usr/bin/env python3
import pandas as pd
def main(filename, min = True, mean = True, max = True):
data = pd.read_csv(filename, index_col=0)
stats = {}
if min:
stats['min'] = data.min(axis=0).min()
if mean:
stats['mean'] = data.mean(axis=0).mean()
if max:
stats['max'] = data.max(axis=0).max()
for key, value in stats.items():
print(f"{key}: {value}")Save it under the name inflammation_stats.py, in ~/Desktop/swc-python/code.
F-strings are a way to format strings in Python. They are defined by prefixing a string with the letter f or F, and they allow you to include expressions inside curly braces {} that will be evaluated at runtime and included in the string. They were introduced in Python 3.6 and provide a more concise and readable way to format strings. See the official documentation for more information on hwo to format strings.
You should already be in the ~/Desktop/swc-python/ directory. If not you can navigate to it using the cd command in the terminal. Then, you can run the script using the following command:
cd ~/Desktop/swc-pythonYou can verify you are in the right directory by running pwd (print working directory):
pwdYou can now run the python script:
./code/inflammation_stats.py It should not output anything, as we only provided a function but did not call it. We need to call the main() function and provide it with the name of the file we want to analyze. We can do that by adding the following lines at the end of our script:
if __name__ == "__main__":
main("data/inflammation-01.csv")This syntax is a common Python idiom that allows us to run the main() function when the script is executed directly from the command line. The if statement check if the script is being run as the main program or if it is being imported as a module in another script. If the script is being run directly, the code inside the if block will be executed. If it is being imported, the code inside the if block will not be executed.
Now, running ./code/inflammation_stats.py in command line should output the average, minimum and maximum inflammation per patient for the file data/inflammation-01.csv.
Running a Python script in bash is very similar to importing that file in Python. The biggest difference is that we don’t expect anything to happen when we import a file, whereas when running a script, we expect to see some output printed to the console.
In order for a Python script to work as expected when imported or when run as a script, we typically put the part of the script that produces output in the following if statement:
if __name__ == '__main__':
main() # Or whatever function produces outputWhen you import a Python file, __name__ is the name of that file (e.g., when importing readings.py, __name__ is inflammation_stats). However, when running a script in bash, __name__ is always set to __main__ in that script so that you can determine if the file is being imported or run as a script.
There are some interesting ways to get input from the user:
input() receives input from the keyboard. This means that the input is defined while the python script is being executed.sys.argv takes arguments provided in command line after the name of the program. This means that the input is defined before the python script is being executed.argparse is similar to sys.argv, with the advantage of being able to give specific names to arguments.The sys.argv list contains the command-line arguments passed to the script, and sys.argv[1] refers to the first argument (the filename in this case).
Python stops executing when it comes to the input() function, and continues when the user has given some input.
In the inflammation_stats.py file , write the following after the end of the main() function:
filename = input("Enter filename: ")
print("Filename is: " + filename)
if __name__ == "__main__":
main(filename)Then in the terminal, run:
./code/inflammation_stats.py You should be asked, in command line, to enter a filename. When you write it (e.g. data/inflammation-01.csv), and press Enter, it should run the main() function with the given file.
Enter filename: data/inflammation-01.csv
Filename is: data/inflammation-01.csv
min: 0
mean: 6.14875
max: 20To use sys.argv you need to import a module called sys. It is part of the standard python library, so you should not have to install anything in particular.
Modify the same python script, to add at the beginning of the script:
import sys and substitute the end of the script with:
print("Filename is: " + sys.argv[1])
if __name__ == "__main__":
main(sys.argv[1])Then in the terminal, run:
./code/inflammation_stats.py data/inflammation-01.csvArguments are given in command line, separated by [space].
Filename is: data/inflammation-01.csv
min: 0
mean: 6.14875
max: 20What is the type of sys.argv? Remember that in python index begins at 0. What do you think is sys.argv[0]? Verify!
Also, what happens if you run ./code/inflammation_stats.py data/inflammation-01.csv data/inflammation-02.csv?
Just like for sys, you need to import argparse.
Modify the same python script, to add at the beginning of the script:
import argparse and substitute the end of the script with:
parser = argparse.ArgumentParser()
parser.add_argument('--filename', action="store", required=True)
args = parser.parse_args()
print("Filename is: " + args.filename)
if __name__ == "__main__":
main(args.filename)Then in the terminal, run:
./code/inflammation_stats.py --filename data/inflammation-01.csvArguments are given in command line, but they have specific names.
argparse is a very useful module when creating programs! You can easily specify the expected type of argument, whether it is optional or not, and create a help for your script. Check their tutorial for more information.
Modify the script to also take --min, --mean and --max as command line arguments. If the argument is given, the corresponding statistic will be calculated and printed. By default, it should not calculate any of the statistics, the user has to specify that in command line.
If the user runs ./code/inflammation_stats_argparse.py --filename data/inflammation-01.csv --min --max, it should only calculate and print the minimum and maximum inflammation per patient, but not the mean.
You can use the action="store_true" option in add_argument() to create a boolean variable for each statistic.
For example:
parser.add_argument('--min', action="store_true")This line will create a variable called args.min, that will be set to False by default. When the user provides the --min argument in command line, args.min will be set to True.
You should successfully run your script with the following commands:
./code/inflammation_stats.py --filename data/inflammation-01.csv --min
./code/inflammation_stats.py --filename data/inflammation-01.csv --max
./code/inflammation_stats.py --filename data/inflammation-01.csv --min --max
./code/inflammation_stats.py --filename data/inflammation-01.csv --min --max --meanThe next step is to teach our program how to handle multiple files.
We want our program to process each file separately, so we need a loop that executes once for each filename. If we specify the files on the command line, the filenames will be in args.filename. Fortunately, argparse has a built-in way to handle an unknown number of arguments. We can use the nargs='+' option in add_argument() to specify that we want to accept one or more filenames as input. This will create a list of filenames in args.filename, which we can then loop through.
Here’s how it can be done:
#!/usr/bin/env python3
import argparse
import pandas as pd
def main(filename, min = True, mean = True, max = True):
for file in filename:
print("Processing file:", file)
data = pd.read_csv(file, index_col=0)
stats = {}
if min:
stats['min'] = data.min(axis=0).min()
if mean:
stats['mean'] = data.mean(axis=0).mean()
if max:
stats['max'] = data.max(axis=0).max()
for key, value in stats.items():
print(f"{key}: {value}")
# Deal with arguments
parser = argparse.ArgumentParser()
parser.add_argument('--filename', nargs='+', action="store", required=True)
parser.add_argument('--min', action="store_true")
parser.add_argument('--max', action="store_true")
parser.add_argument('--mean', action="store_true")
args = parser.parse_args()
if __name__ == "__main__":
main(args.filename, min=args.min, mean=args.mean, max=args.max)Then in command line, you can run:
./code/inflammation_stats.py --filename data/inflammation-01.csv --min --max
./code/inflammation_stats.py --filename data/inflammation-01.csv data/inflammation-02.csv --min --max
./code/inflammation_stats.py --filename data/inflammation-0*.csv --min --maxWhat is the type of args.filename when using nargs='+'? What does it contain?
When coding, a good practice is to test your code on small data files before running it on large datasets. This allows you to check that your code is working as expected and to catch any errors or bugs before they cause problems with larger datasets.
Indeed, your code might not have any syntax errors, but it might not be doing what you think it is doing. For example, you might think that your code is calculating the mean of a dataset, but in reality, it is calculating the sum. If you run your code on a large dataset, you might not notice this error until it is too late.
Here we can test our code on a smaller file that has the same structure as the larger files, but only contains a few lines of data. This way, we can easily check that our code is calculating the mean correctly for each line, and that it is doing what we expect it to do.
The file data/small-01.csv contains the following data:
Day1,Day2,Day3
Patient1,0,0,1
Patient2,0,1,2We can run:
./code/inflammation_stats.py --filename data/small-01.csv --min --max --meanThere are two more files small-02.csv and small-03.csv that you can use to test your code.
It is also important to test your code on edge cases, which are cases that are at the extreme ends of the input space. For example, if your code is supposed to handle missing values, you should test it on a file that contains missing values to make sure that it is handling them correctly.
Even if your documentation is very clear, the user might not behave as you expect them to, and might provide input that is not in the format you expected. For example, if your code is supposed to take a filename as input, the user might provide a filename that does not exist, or a filename that is in a different format than what you expected. It is important to test your code on such edge cases to make sure that it is handling them correctly and providing useful error messages to the user.
Every programmer encounters errors, both those who are just beginning, and those who have been programming for years. Encountering errors and exceptions can be very frustrating at times, and can make coding feel like a hopeless endeavour. However, understanding what the different types of errors are and when you are likely to encounter them can help a lot. Once you know why you get certain types of errors, they become much easier to fix.
Errors in Python have a very specific form, called a traceback. Let’s examine one by providing a filename that does not exist:
./code/inflammation_stats.py --filename data/small-06.csv --min --max --meanProcessing file: data/small-06.csv
Traceback (most recent call last):
File "/Volumes/projects/PCa_SingleCell/utils/workshop/python-intro/./code/inflammation_stats.py", line 31, in <module>
main(args.filename, min=args.min, mean=args.mean, max=args.max)
File "/Volumes/projects/PCa_SingleCell/utils/workshop/python-intro/./code/inflammation_stats.py", line 9, in main
data = pd.read_csv(file, index_col=0)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/gilbartv/miniconda3/lib/python3.11/site-packages/pandas/io/parsers/readers.py", line 1026, in read_csv
return _read(filepath_or_buffer, kwds)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/gilbartv/miniconda3/lib/python3.11/site-packages/pandas/io/parsers/readers.py", line 620, in _read
parser = TextFileReader(filepath_or_buffer, **kwds)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/gilbartv/miniconda3/lib/python3.11/site-packages/pandas/io/parsers/readers.py", line 1620, in __init__
self._engine = self._make_engine(f, self.engine)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/gilbartv/miniconda3/lib/python3.11/site-packages/pandas/io/parsers/readers.py", line 1880, in _make_engine
self.handles = get_handle(
^^^^^^^^^^^
File "/Users/gilbartv/miniconda3/lib/python3.11/site-packages/pandas/io/common.py", line 873, in get_handle
handle = open(
^^^^^
FileNotFoundError: [Errno 2] No such file or directory: 'data/small-06.csv'Python tells us the category or type of error (in this case, it is an FileNotFoundError) and a more detailed error message (in this case, it says [Errno 2] No such file or directory: 'data/small-06.csv').
This particular traceback has many levels. The traceback also shows us the exact line of code where the error occurred, and the sequence of function calls that led to that line of code being executed.
The lines in our code that caused the error are main(args.filename, min=args.min, mean=args.mean, max=args.max), and then pd.read_csv(file, index_col=0). Indeed main() is the function that we called in command line, and inside main(), the error occurred when trying to read the file with pd.read_csv(). It gives more or the subsequent calls that were made inside the pandas library to get to the line of code that caused the error, but we can ignore those for now.
The length of the error message does not reflect severity, rather, it indicates that your program called many subsequent functions before it encountered the error.
One reason for receiving this FileNotFoundError error is that you specified an incorrect path to the file. For example, if I am currently in a folder called myproject, and I have a file in myproject/writing/myfile.txt, but I try to open myfile.txt, this will fail. The correct path would be writing/myfile.txt.
It is also possible that the file name or its path contains a typo.
If you encounter an error and don’t know what it means, it is still important to read the traceback closely. That way, if you fix the error, but encounter a new one, you can tell that the error changed. Additionally, sometimes knowing where the error occurred is enough to fix it, even if you don’t entirely understand the message.
If you do encounter an error you don’t recognize, try looking at the official documentation on errors also called Exceptions in Python. However, note that you may not always be able to find the error there, as it is possible to create custom errors. In that case, hopefully the custom error message is informative enough to help you figure out what went wrong.
Here is a table of some of the built-in exceptions in python.
| Exception | Description |
|---|---|
IndexError |
Raised when the index of a sequence is out of range. |
KeyError |
Raised when a key is not found in a dictionary. |
KeyboardInterrupt |
Raised when the user hits the interrupt key (Ctrl+c or Delete). |
NameError |
Raised when a variable is not found in the local or global scope. |
TypeError |
Raised when a function or operation is applied to an object of an incorrect type. |
StatementError |
Raised when statements are in an order or contain characters not expected by the programming language. |
ValueError |
Raised when a function receives an argument of the correct type but of an incorrect value. |
RuntimeError |
Raised when an error occurs that do not belong to any specific exceptions. |
Exception |
Base class of exceptions. |
Try the following code and understand what the error is:
# Code A
def print_message(day):
messages = [
'Hello, world!',
'Today is Tuesday!',
'It is the middle of the week.',
'Today is Donnerstag in German!',
'Last day of the week!',
'Hooray for the weekend!',
'Aw, the weekend is almost over.'
]
print(messages[day])
def print_sunday_message():
print_message(7)
print_sunday_message()--------------------------------------------------------------------------- IndexError Traceback (most recent call last) Cell In[57], line 17 14 def print_sunday_message(): 15 print_message(7) ---> 17 print_sunday_message() Cell In[57], line 15, in print_sunday_message() 14 def print_sunday_message(): ---> 15 print_message(7) Cell In[57], line 12, in print_message(day) 2 def print_message(day): 3 messages = [ 4 'Hello, world!', 5 'Today is Tuesday!', (...) 10 'Aw, the weekend is almost over.' 11 ] ---> 12 print(messages[day]) IndexError: list index out of range
# Code B
def some_function()
msg = 'hello, world!'
print(msg)
return msgCell In[58], line 2 def some_function() ^ SyntaxError: expected ':'
# Code C
def some_function():
msg = 'hello, world!'
print(msg)
return msgCell In[59], line 5 return msg ^ TabError: inconsistent use of tabs and spaces in indentation
# Code D
print(a)--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[60], line 2 1 # Code D ----> 2 print(a) NameError: name 'a' is not defined
# Code E
for number in range(10):
total = total + number
print('The count is:', count)--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[61], line 3 1 # Code E 2 for number in range(10): ----> 3 total = total + number 4 print('The count is:', count) NameError: name 'total' is not defined
# Code F
Count = 0
for number in range(10):
count = count + number
print('The count is:', count)--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[62], line 4 2 Count = 0 3 for number in range(10): ----> 4 count = count + number 5 print('The count is:', count) NameError: name 'count' is not defined
It is possible to handle errors (in python, they are also called exceptions), using the following statements:
try to test a block of code for errorsexcept to handle the errorelse to excute code if there is no errorfinally to excute code, regardless of the result of the try and except blocks# The try block will generate an exception, because some_undefined_variable is not defined:
try:
print(some_undefined_variable)
except:
print("Oops... Something went wrong") Oops... Something went wrong# Without the try block, the program will crash and raise an error:
print(some_undefined_variable)--------------------------------------------------------------------------- NameError Traceback (most recent call last) Cell In[64], line 2 1 # Without the try block, the program will crash and raise an error: ----> 2 print(some_undefined_variable) NameError: name 'some_undefined_variable' is not defined
try:
print(some_undefined_variable)
except:
print("Oops... Something went wrong")
else:
print("Nothing went wrong")
finally:
print("The 'try except' is finished") Oops... Something went wrong
The 'try except' is finishedYou can use the Exceptions types to be more specific about the type of exception occurring.
try:
print(some_undefined_variable)
except NameError:
print("A variable is not defined")
except:
print("Oops... Something went wrong")
else:
print("Nothing went wrong")
finally:
print("The 'try except' is finished") A variable is not defined
The 'try except' is finishedYou can also use the Exceptions types to throw an exception if a condition occurs, by using the raise keyword.
x = "hello"
try:
if not isinstance(x, int):
raise TypeError("Only integers are allowed")
if x < 0:
raise ValueError("Sorry, no numbers below zero")
print(x, "is a positive integer.")
except NameError:
print("A variable is not defined")
else:
print("Nothing went wrong")
finally:
print("The 'try except' is finished") The 'try except' is finished--------------------------------------------------------------------------- TypeError Traceback (most recent call last) Cell In[67], line 4 2 try: 3 if not isinstance(x, int): ----> 4 raise TypeError("Only integers are allowed") 5 if x < 0: 6 raise ValueError("Sorry, no numbers below zero") TypeError: Only integers are allowed
Let’s make our previous function even better by adding some exception handling. Raise a TypeError if the input filename is not a string. To verify that a variable is of a certain type, you can use the isinstance() function. For example, isinstance(x, str) will return True if x is a string, and False otherwise.
With the input given below, the output and errors should be:
import pandas as pd
def main(filename, min = True, mean = True, max = True):
print("Processing file:", filename)
data = pd.read_csv(filename, index_col=0)
stats = {}
if min:
stats['min'] = data.min(axis=0).min()
if mean:
stats['mean'] = data.mean(axis=0).mean()
if max:
stats['max'] = data.max(axis=0).max()
for key, value in stats.items():
print(f"{key}: {value}")
main(filename = 5474)
main(filename = "data/inflammation-01.csv")--------------------------------------------------------------------------- TypeError Traceback (most recent call last) Cell In[68], line 18 15 for key, value in stats.items(): 16 print(f"{key}: {value}") ---> 18 main(filename = 5474) 19 main(filename = "data/inflammation-01.csv") Cell In[68], line 5, in main(filename, min, mean, max) 3 def main(filename, min = True, mean = True, max = True): 4 if not isinstance(filename, str): ----> 5 raise TypeError(f"Filename must be a string, {filename} is of type {type(filename)}") 6 print("Processing file:", filename) 7 data = pd.read_csv(filename, index_col=0) TypeError: Filename must be a string, 5474 is of type <class 'int'>
The first step in debugging something is to know what it’s supposed to do. “My program doesn’t work” isn’t good enough: in order to diagnose and fix problems, we need to be able to tell correct output from incorrect. If we can write a test case for the failing case — i.e., if we can assert that with these inputs, the function should produce that result — then we’re ready to start debugging. If we can’t, then we need to figure out how we’re going to know when we’ve fixed things.
But writing test cases for scientific software is frequently harder than writing test cases for commercial applications, because if we knew what the output of the scientific code was supposed to be, we wouldn’t be running the software: we’d be writing up our results and moving on to the next program. In practice, scientists tend to do the following:
We can only debug something when it fails, so the second step is always to find a test case that makes it fail every time. The “every time” part is important because few things are more frustrating than debugging an intermittent problem: if we have to call a function a dozen times to get a single failure, the odds are good that we’ll scroll past the failure when it actually occurs.
As part of this, it’s always important to check that our code is “plugged in”, i.e., that we’re actually exercising the problem that we think we are. Every programmer has spent hours chasing a bug, only to realize that they were actually calling their code on the wrong data set or with the wrong configuration parameters, or are using the wrong version of the software entirely. Mistakes like these are particularly likely to happen when we’re tired, frustrated, and up against a deadline, which is one of the reasons late-night (or overnight) coding sessions are almost never worthwhile.
If it takes 20 minutes for the bug to surface, we can only do three experiments an hour. This means that we’ll get less data in more time and that we’re more likely to be distracted by other things as we wait for our program to fail, which means the time we are spending on the problem is less focused. It’s therefore critical to make it fail fast.
As well as making the program fail fast in time, we want to make it fail fast in space, i.e., we want to localize the failure to the smallest possible region of code:
The smaller the gap between cause and effect, the easier the connection is to find. Many programmers therefore use a divide and conquer strategy to find bugs, i.e., if the output of a function is wrong, they check whether things are OK in the middle, then concentrate on either the first or second half, and so on.
N things can interact in N! different ways, so every line of code that isn’t run as part of a test means more than one thing we don’t need to worry about.
Replacing random chunks of code is unlikely to do much good. (After all, if you got it wrong the first time, you’ll probably get it wrong the second and third as well.) Good programmers therefore change one thing at a time, for a reason. They are either trying to gather more information (“is the bug still there if we change the order of the loops?”) or test a fix (“can we make the bug go away by sorting our data before processing it?”).
Every time we make a change, however small, we should re-run our tests immediately, because the more things we change at once, the harder it is to know what’s responsible for what (those N! interactions again). And we should re-run all of our tests: more than half of fixes made to code introduce (or re-introduce) bugs, so re-running all of our tests tells us whether we have regressed.
Good scientists keep track of what they’ve done so that they can reproduce their work, and so that they don’t waste time repeating the same experiments or running ones whose results won’t be interesting. Similarly, debugging works best when we keep track of what we’ve done and how well it worked. If we find ourselves asking, “Did left followed by right with an odd number of lines cause the crash? Or was it right followed by left? Or was I using an even number of lines?” then it’s time to step away from the computer, take a deep breath, and start working more systematically.
Records are particularly useful when the time comes to ask for help. People are more likely to listen to us when we can explain clearly what we did, and we’re better able to give them the information they need to be useful.
And speaking of help: if we can’t find a bug in 10 minutes, we should be humble and ask for help. Explaining the problem to someone else is often useful, since hearing what we’re thinking helps us spot inconsistencies and hidden assumptions. If you don’t have someone nearby to share your problem description with, get a rubber duck!
Asking for help also helps alleviate confirmation bias. If we have just spent an hour writing a complicated program, we want it to work, so we’re likely to keep telling ourselves why it should, rather than searching for the reason it doesn’t. People who aren’t emotionally invested in the code can be more objective, which is why they’re often able to spot the simple mistakes we have overlooked.
Part of being humble is learning from our mistakes. Programmers tend to get the same things wrong over and over: either they don’t understand the language and libraries they’re working with, or their model of how things work is wrong. In either case, taking note of why the error occurred and checking for it next time quickly turns into not making the mistake at all.
If we train ourselves to avoid making some kinds of mistakes, to break our code into modular, testable chunks, and to turn every assumption (or mistake) into an assertion, it will actually take us less time to produce working programs, not more.
Here are a couple of tips:
You can follow some free tutorials on:
Here are some references and ressources that greatly inspired this class :